James Collins, Ph.D.
AIMBE College of Fellows Class of 2000 For innovative combinations of nonlinear biodynamics and biomechanics to advanced understanding in and applications to posture control and neurosensory function.
Scientists discover compounds that help cells fight a wide range of viruses
Via MIT | July 14, 2025The molecules trigger a built-in cellular stress response and show promise as broad-spectrum antivirals against Zika, herpes, RSV, and more.
Researchers at MIT and other institutions have identified compounds that can fight off viral infection by activating a defense pathway inside host cells. These compounds, they believe, could be used as antiviral drugs that work against not just one but any kind of virus.
The researchers identified these compounds, which activate a host cell defense system known as the integrated stress response pathway, in a screen of nearly 400,000 molecules. In tests in human cells, the researchers showed that the compounds help cells fend off infection from RSV, herpes virus, and Zika virus. They also proved effective in combating herpes infection in a mouse model… Continue reading.
Using AI, MIT researchers identify a new class of antibiotic candidates
Via MIT | December 20, 2023These compounds can kill methicillin-resistant Staphylococcus aureus (MRSA), a bacterium that causes deadly infections.
Using a type of artificial intelligence known as deep learning, MIT researchers have discovered a class of compounds that can kill a drug-resistant bacterium that causes more than 10,000 deaths in the United States every year.
In a study appearing today in Nature, the researchers showed that these compounds could kill methicillin-resistant Staphylococcus aureus (MRSA) grown in a lab dish and in two mouse models of MRSA infection. The compounds also show very low toxicity against human cells, making them particularly good drug candidates… Continue reading.
A new control switch could make RNA therapies easier to program
Via MIT | March 15, 2023Using this approach, researchers hope to deliver therapeutic RNA molecules selectively to cancer cells or other target cells.
Using an RNA sensor, MIT engineers have designed a new way to trigger cells to turn on a synthetic gene. Their approach could make it possible to create targeted therapies for cancer and other diseases, by ensuring that synthetic genes are activated only in specific cells.
The researchers demonstrated that their sensor could accurately identify cells expressing a mutated version of the p53 gene, which drives cancer development, and turn on a gene encoding a fluorescent protein only within those cells. In future work, they plan to develop sensors that would trigger production of cell-killing proteins in cancer cells, while sparing healthy cells… Continue reading.
Analyzing the potential of AlphaFold in drug discovery
Via MIT | September 6, 2022Study finds computer models that predict molecular interactions need improvement before they can help identify drug mechanisms of action.
Over the past few decades, very few new antibiotics have been developed, largely because current methods for screening potential drugs are prohibitively expensive and time-consuming. One promising new strategy is to use computational models, which offer a potentially faster and cheaper way to identify new drugs.
A new study from MIT reveals the potential and limitations of one such computational approach. Using protein structures generated by an artificial intelligence program called AlphaFold, the researchers explored whether existing models could accurately predict the interactions between bacterial proteins and antibacterial compounds. If so, then researchers could begin to use this type of modeling to do large-scale screens for new compounds that target previously untargeted proteins. This would enable the development of antibiotics with unprecedented mechanisms of action, a task essential to addressing the antibiotic resistance crisis… Continue reading.
Engineered bacteria could help protect “good” gut microbes from antibiotics
Via MIT | April 11, 2022Antibiotics are life-saving drugs, but they can also harm the beneficial microbes that live in the human gut. Following antibiotic treatment, some patients are at risk of developing inflammation or opportunistic infections such as Clostridiodes difficile. Indiscriminate use of antibiotics on gut microbes can also contribute to the spread of resistance to the drugs.
In an effort to reduce those risks, MIT engineers have developed a new way to help protect the natural flora of the human digestive tract. They took a strain of bacteria that is safe for human consumption and engineered it to safely produce an enzyme that breaks down a class of antibiotics called beta-lactams. These include ampicillin, amoxicillin, and other commonly used drugs… Continue reading.
Engineers devise a way to selectively turn on RNA therapies in human cells
Via MIT | October 28, 2021Researchers at MIT and Harvard University have designed a way to selectively turn on gene therapies in target cells, including human cells. Their technology can detect specific messenger RNA sequences in cells, and that detection then triggers production of a specific protein from a transgene, or artificial gene.
Because transgenes can have negative and even dangerous effects when expressed in the wrong cells, the researchers wanted to find a way to reduce off-target effects from gene therapies. One way of distinguishing different types of cells is by reading the RNA sequences inside them, which differ from tissue to tissue… Continue reading.
New face mask prototype can detect Covid-19 infection
Via MIT | June 28, 2021Engineers at MIT and Harvard University have designed a novel face mask that can diagnose the wearer with Covid-19 within about 90 minutes. The masks are embedded with tiny, disposable sensors that can be fitted into other face masks and could also be adapted to detect other viruses.
The sensors are based on freeze-dried cellular machinery that the research team has previously developed for use in paper diagnostics for viruses such as Ebola and Zika. In a new study, the researchers showed that the sensors could be incorporated into not only face masks but also clothing such as lab coats, potentially offering a new way to monitor health care workers’ exposure to a variety of pathogens or other threats… Continue reading.
Wyss Institute’s nasal swab and toehold switch technologies licensed to facilitate SARS-CoV-2 diagnostic efforts
Via Harvard University | October 5, 2020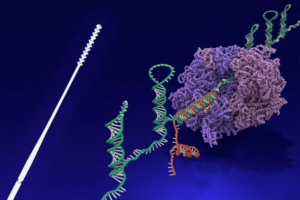 …
…
Toehold switches could come into play at the other end of the COVID-19 diagnostic process. Pioneered in the groups of Wyss Core Faculty members James Collins, Ph.D., and Peng Yin, Ph.D., they are synthetic nucleic acid-based devices that function as sensors for external stimuli (“inputs”), like RNA molecules derived from pathogenic viruses. When integrated into synthetic gene circuits, Toehold Switches can be designed to turn on a gene of interest, which can be a reporter signaling the presence of the environmental stimulus. In their OFF state, these nanotechnological devices form a hairpin-like structure that specifically associates with and actively blocks the expression of a (reporter) target gene. Once an “input” RNA binds to their “toehold” region, the hairpin structure opens up and adopts an ON state to allow the protein-synthesizing machinery access to the target gene, which results in the synthesis of the actual signaling molecule.
In a series of proof-of-concept studies, the Collins and Yin teams have demonstrated toehold switches to function in living cells as computational devices that can assess and report complex combinations of environmental stimuli. They also have utilized them as key components of paper-based synthetic gene circuits that can be applied as diagnostics to sense and indicate different pathogens, including Ebola and Zika viruses. The versatile capabilities of Toehold Switches offer an opportunity for inexpensively and effectively surveilling the presence of pathogens with high sensitivity and specificity in different environmental settings, including working environments during the reopening phase of the pandemic… Continue reading.
Artificial intelligence helps researchers find new antibiotics
Via Science Board | March 31, 2020To address antibiotic resistance, researchers have developed a machine-learning approach that can search millions of known chemicals to find potential new antimicrobial compounds. This research, published in Cell on February 20, uncovered several promising antibiotic candidates that will move into clinical testing.
After training a deep neural network to identify potential antibiotics that kill bacteria using different mechanisms than current drugs, a team of researchers led by James Collins, PhD, from Massachusetts Institute of Technology (MIT) discovered a new antibiotic compound that killed many challenging pathogens… Continue reading.
Biomaterials Smarten up with CRISPR
Via Lab Manager | August 23, 2019The CRISPR-Cas system has become the go-to tool for researchers who study genes in an ever-growing list of organisms, and is being used to develop new gene therapies that potentially can correct a defect at a single nucleotide position of the vast reaches of the genome. It is also being harnessed in ongoing diagnostic approaches for the detection of pathogens and disease-causing mutations in patients.
Now, reporting in Science, a research team at Harvard’s Wyss Institute for Biologically Inspired Engineering and the Massachusetts Institute of Technology (MIT) demonstrates the use of CRISPR as a control element in a new type of stimuli-responsive “smart” materials. Upon activation by specific natural or user-defined DNA stimuli, a CRISPR-Cas enzyme enables a variety of smart materials to release bound cargo such as fluorescent dyes and active enzymes, change their structures to deploy encapsulated nanoparticles and live cells, or regulate electric circuits thereby converting biological into electric signals… Continue reading.
Inexpensive biology kits offer hands-on experience with DNA
Via MIT | August 1, 2018To help students gain a better grasp of biological concepts, MIT and Northwestern University researchers have designed educational kits that can be used to perform experiments with DNA, to produce glowing proteins, scents, or other easily observed phenomena.
Biology teachers could use the BioBits kits to demonstrate key concepts such as how DNA is translated into proteins, or students could use them to design their own synthetic biology circuits, the researchers say.
“Our vision is that these kits will serve as a creative outlet for young individuals, and show them that biology can be a design platform,” says James Collins, the Termeer Professor of Medical Engineering and Science in MIT’s Institute for Medical Engineering and Science (IMES) and Department of Biological Engineering. “The time is right for creating educational kits that could be utilized in classrooms or in the home, to introduce young folks as well as adults who want to be retrained in biotech, to the technologies that underpin synthetic biology and biotechnology… Continue reading.
Boosting the antibiotic arsenal
Via MIT | December 7, 2017MIT researchers have discovered a way to make bacteria more vulnerable to a class of antibiotics known as quinolones, which include ciprofloxacin and are often used to treat infections such as Escherichia coli and Staphylococcus aureus.
The new strategy overcomes a key limitation of these drugs, which is that they often fail against infections that feature a very high density of bacteria. These include many chronic, difficult-to-treat infections, such as Pseudomonas aeruginosa, often found in the lungs of cystic fibrosis patients, and methicillin-resistant Staphylococcus aureus (MRSA).
“Given that the number of new antibiotics being developed is diminishing, we face challenges in treating these infections. So efforts such as this could enable us to expand the efficacy of existing antibiotics,” says James Collins, the Termeer Professor of Medical Engineering and Science in MIT’s Institute for Medical Engineering and Science (IMES) and Department of Biological Engineering and the senior author of the study… Continue reading.
Finding Zika One Paper Disc at a Time
Via PHYS.org | May 6, 2016An international, multi-institutional team of researchers led by synthetic biologist James Collins, Ph.D. at the Wyss Institute for Biologically Inspired Engineering at Harvard University, has developed a low-cost, rapid paper-based diagnostic system for strain-specific detection of the Zika virus, with the goal that it could soon be used in the field to screen blood, urine, or saliva samples.
“The growing global health crisis caused by the Zika virus propelled us to leverage novel technologies we have developed in the lab and use them to create a workflow that could diagnose a patient with Zika, in the field, within 2-3 hours,” said Collins, who is a Wyss Core Faculty member, and Termeer Professor of Medical Engineering & Science and Professor of Biological Engineering at the Massachusetts Institute of Technology (MIT)’s Department of Biological Engineering.
Human-Gut-On-A-Chip Model Offers Hope For IBD Sufferers
Via Harvard | December 15, 2015It’s estimated that as many as a million Americans suffer from inflammatory bowel diseases (IBD) such as ulcerative colitis and Crohn’s disease, which cause mild to severe symptoms that at best can be managed and at worst lead to life-threatening complications.
While abnormal immune responses are largely responsible for these diseases, issues relating to gut microbiome, intestinal epithelial cells, immune components, and the gut’s rhythmic peristalsis motions can also contribute to and exacerbate symptoms. But until now, scientists have been hard pressed to develop new therapies for treating IBDs because they could not replicate the human gut microenvironment in the laboratory.
On Monday, the Wyss Institute for Biologically Inspired Engineering at Harvard University announced that its team had created a model of human intestinal inflammation and bacterial overgrowth in a human-gut-on-a-chip. The team, co-led by Wyss Institute Founding Director Donald Ingber and core faculty member James Collins, leveraged the institute’s proprietary human-organs-on-chips technology to microengineer the model.
“There is much to be learned about IBD, as well as how antibiotics impact the microbiome,” said Collins, the Termeer Professor of Bioengineering in the Department of Biological Engineering and Institute for Medical Engineering & Science at the Massachusetts Institute of Technology. “This technology enables one to study in an isolated and controlled manner the complexity of the microbiome and the role different microbial species play in health and disease. It is therefore a highly valuable platform for discovery and clinical translation efforts.”
From Dance Clubs to Syn Bio: MIT’s Collins on Startups, Second Chances
Via Xconomy | October 5, 2015t happens over and over again with new science. A discovery prompts crazy hype and massive investment that the data aren’t ready to support. A crash ensues, backers lose millions, egos are bruised—yet the pioneers slowly trudge forward. They regroup, away from the limelight, and try to learn from failure.
When it comes to synthetic biology—a method of modifying the genes of living organisms to effectively change what they do—James Collins knows this story better than most. He’s an MIT professor who helped found the field nearly two decades ago. He’s seen the hype, when investors placed huge bets on startups aiming to produce clean energy on a large scale; the crash, when many of those companies were wiped out and scientists fled back to academia; and the pivot, when the surviving companies shifted their sights elsewhere.
“I think we’ve recovered now, as a field,” he says.
Gone are the days when a bevy of high-profile startups like Sapphire Energy, Solazyme, and J. Craig Venter’s LS9 offered hopes of renewable, eco-friendly fuels made by engineered algae. In their wake is diversification: Sapphire, for instance, has made a strategic pivot into things like food additives, cosmetics, and nutraceuticals. But from Collins’s vantage point, something else has happened. The “clinical space,” he says, has become a dominant focus for synthetic biologists—meaning tools that could be used for medical research, diagnostics, or even “living” therapeutics like the ones Cambridge, MA-based Synlogic, a startup from Collins’s lab, is trying to develop.
Collins (pictured above) is a New York-New England hybrid. He was born in the Bronx before moving first to Bellerose, in the outskirts of Queens, and later, after he finished elementary school, to New Hampshire. He used to have a strong New York accent and, as a Queens guy, was a fan of the Jets, Mets, and Nets. (Former Nets small forward William “Billy” Schaeffer, who also grew up in Bellerose, would shoot hoops nearby.) Now that Big Apple accent is largely gone (“I joke that I’ve got a New York attitude but not a New York accent,” he says) and Collins shows a fierce allegiance to all teams Boston. He even threw out the first pitch at a Red Sox game in 2008 at Fenway Park.
Five Takeaways From “Boston’s Life Science Disruptors”
Via Xconomy | October 2, 2015—Cool technology. Now, what to we do with it? Atlas Venture’s Peter Barrett and Ankit Mahadevia were interested in MIT professor Jim Collins and protégé Timothy Lu’s latest work. The synthetic biology specialists had two things cooking: a tools platform to “rewire organisms,” and an idea for engineered microbes that could serve as living drugs or diagnostics. Mahadevia’s response: “We really want to do something with you guys, we just don’t know what.”
A month later, Atlas decided no to the tools, and yes to the living drugs. But how to use them? Collins’s preference—wiping out infectious diseases, like a cholera infection. Mahadevia’s response: “We still want to do something with you guys, we just don’t know what.”
It took a suggestion from Atlas entrepreneur-in-residence Dean Falb to figure out what to do. He showed Collins a list of 12 rare genetic metabolic disorders, all of which Collins had never heard of. The microbes made sense here—they could either make a metabolite that is missing, or break down a toxic one. The medical need is significant, and Synlogic could run small, inexpensive clinical trials for it.
“I thought this was brilliant,” Collins said. Synlogic is now targeting phenylketonuria and urea cycle disorders.
Building a Paper Gene Circuit
Via BU Engineering | December 1, 2014The first case of the Ebola outbreak currently ravaging West Africa appeared in Guinea in December 2013. But it wasn’t until March 22, 2014, that scientists finally confirmed the virus as Ebola. By that point, 49 people had already died.
Why did it take so long? Partly because confirming the diagnosis required that epidemiologists fly from Europe to Africa, collect blood samples, fly back to Europe, and analyze them in sophisticated labs.
Now a team of biologists at BU, led by Professor James Collins (BME, MSE, SE), has created a new tool that could provide a quick, cheap way to perform sophisticated lab analyses and diagnostics in the field, and may also offer a way to speed science in the lab. The tool, called a paper gene circuit, takes biological reactions out of cells and puts them onto a piece of paper. It is described in the November 6, 2014, issue of Cell.
“This could really be a game-changer for a lot of applications, including diagnostics,” says Collins, who is also a core faculty member at Harvard’s Wyss Institute. “You can literally carry this in your pocket and run an experiment in the field without any additional equipment.”
Beyond Bacteria
Via Boston University | July 7, 2014At the heart of synthetic biology is the assembly of genetic components into “circuits” that perform desired operations in living cells, with the long-term goal of empowering these cells to solve critical problems in healthcare, energy, the environment and other domains, from cancer treatment to toxic waste cleanup. While much of this work is done using bacterial cells, new techniques are emerging to reprogram eukaryotic cells—those found in plants and animals, including humans—to perform such tasks.
To engineer useful genetic circuits in eukaryotic cells, synthetic biologists typically manipulate sequences of DNA in an organism’s genome, but Assistant Professor Ahmad “Mo” Khalil (BME), Professor James J. Collins (BME, MSE, SE) postdoctoral fellow Albert J. Keung (BME) and other researchers at Boston University’s Center of Synthetic Biology (CoSBi) have another idea that could vastly increase their capabilities. Rather than manipulate the DNA sequence directly, the CoSBi engineers are exploiting a class of proteins that regulate chromatin, the intricate structure of DNA and proteins that condenses and packages a genome to fit within the cell. These chromatin regulator (CR) proteins play a key role in expressing—turning on and off—genes throughout the cell, so altering their makeup could provide a new pathway for engineering the cell’s genetic circuits to perform desired functions.
Collins Elected to National Academy of Sciences: Achieves “Trifecta”
Via BU Biomedical Engineering | May 1, 2014Professor James J. Collins (BME, MSE, SE) has been elected to the National Academy of Sciences(NAS), one of the highest honors in science and technology, in recognition of his distinguished and continuing achievements in original research.
Collins, who is one of the founders of the field of synthetic biology, joins Boston University’s seven other NAS members, a group that includes President Robert A. Brown, Nobel Prize–winning theoretical physicist Sheldon Glashow, BU’s Arthur G. B. Metcalf Professor of Mathematics and Science, and Nancy Kopell, a William Fairfield Warren Distinguished Professor. The NAS, a private, nonprofit society of distinguished scholars, is charged with providing independent, objective advice to the nation on science and technology. Collins is one of 84 new members and 21 foreign associates.
…
“We are thrilled to learn of Jim’s election to the National Academy of Sciences,” said Jean Morrison, University provost and chief academic officer. “This is one of the most significant honors for a scientist, and Jim is well deserving of this recognition. Jim’s pathbreaking research in synthetic and systems biology, with a particular focus on antibiotics, has set him apart as one of the world’s top researchers—and we are extraordinarily proud to have him as a member of the BU faculty.”
 AIMBE
AIMBE
